|
[ Up ] [ Rotavirus figure ]
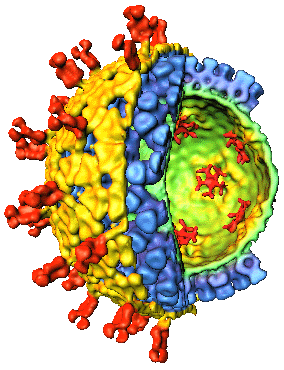
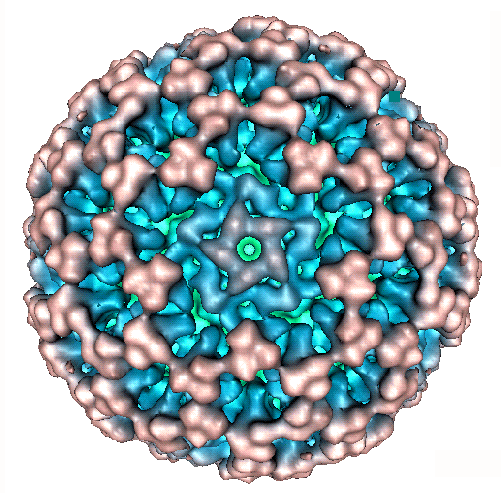
The dsRNA segments
and proteins of simian rotavirus A / SA11
(genus Rotavirus:
family Reoviridae)
(available as a Word
file )
Coding
assignments, virion locations of rotavirus proteins and
3D structure.
Structural
studies [17, 31, 37, 72, 73, 88, 89]: last updated April 2003, references in
[square brackets]
|
Genome
Segment†
Size
(bp) |
ORFs |
Gene
Product(s)
( ':
Protein Function) |
Protein
Size |
Location
in Virus Particle |
Copy
Number/
Particle |
Cognate
Proteins‡ |
GenBank
Accession Number (s) |
Functions
and Properties |
|
1
(3302) |
18-3282 |
VP1
(Pol) |
1088
aa
125,005
Da |
Inner
capsid, 5-fold axis |
12 |
Orthoreovirus l3(Pol)
Orbivirus VP
1(Pol)
Coltivirus
VP1(Pol)
Cypovirus Pol |
X16830
[56] |
RNA-dependent
RNA polymerase [87].
Part of minimal
replication complex [63,87],
Virus specific
3’-mRNA binding [61,62]
Part of virion
transcription complex with VP3 [11,73] |
|
2
(2690) |
17-2659 |
VP2
(T1) |
881
aa
102,431
Da |
Inner
capsid |
120 |
Orbivirus:
VP3
Orthoreovirus:
l1 |
X16831
[56] |
Inner
capsid structural protein [8].
Non-specific ss
& dsRNA-binding activity [10]
Myristoylated
[15].
Cleaved
[23,86].
Part of minimal
replication complex [63].
Leucine zipper
[56].
Interacts
with VP5 [7]. |
|
3
(2591) |
50-2554 |
VP3
(Cap) |
835
aa
98,120
Da |
Inner
capsid, 5-fold axis |
12 |
Orbivirus:
VP4
Orthoreovirus:
l2 |
X16062
[44]
X16387
[56] |
Guanylyltransferase
[45,68].
Methyltransferase
[13].
Basic Protein
[44,56].
Part of virion
transcription complex with VP1 [11,73].
Non-specific
ssRNA binding [62]. |
|
4
(2362) |
10
-2337 |
VP4 |
776
aa
86,782 |
Outer
capsid spike |
120 |
- |
D16346
[77]
X14204
[55] |
VP4 Dimers form
outer capsid spike [3].
Interacts with
VP6 [89].
Virus
infectivity enhanced by trypsin cleavage of VP4 into VP5* and VP8*
[22,46].
Hemagglutinin
[26,38].
Cell attachment
protein [47,75,85].
P-type
neutralization antigen [32,58].
VP5*
permeabilizes membranes [16].
Crystal
structure of VP8 fragment (galectin fold) [19].
TRAF2 signaling
[43].
Protection
[33]. |
|
- |
VP5* |
529
aa
(247-776)
60,000
Da |
|
|
|
- |
VP8* |
247
aa
(1-247) 28,000
Da |
|
|
|
5
(1611) |
31-1515 |
NSP1 |
495
aa
58,654
Da |
Nonstructural |
0 |
- |
L18944
[35]
X14914
[57] |
Associates with
cytoskeleton [34].
Extensive
sequence diversity between strains [20,42,57].
Two conserved
cysteine-rich zinc-finger motifs [57,60].
Virus specific
5’-mRNA binding [34,62].
Interacts with
host IFN regulatory factor 3 [29]. |
|
6
[1356] |
24-1214 |
VP6
(T13) |
397
aa
48,160
Da |
Middle
capsid |
780 |
Orbivirus:
VP7 |
L15384
[48]
L33365
[48]
M27824
[76] |
Major virion
protein [49,72].
Middle capsid
structural protein [72].
Homotrimeric 4o
structure [72].
Subgroup
antigen [30,39].
Myristoylated
[15].
Protection (?
Mechanism) [11,84].
Crystal
structure [50].
Hydrophobic
[48,76]. |
|
7
(1105) |
26-970 |
NSP3 |
315
aa
34,600 Da |
Nonstructural |
0 |
- |
M87502
[51] |
Homodimer
[51,66].
Virus-specific
3’- mRNA binding [69,70].
Binds eIF4G1
and circularizes mRNA on initiation complex [67].
Involved in
translational regulation and host shut-off [14,59,82].
Crystal
structure: NSP3 NH3 fragment with 3’- viral RNA [17] and
NSP3 COOH fragment with eIF4G fragment [31]. |
|
8
(1059) |
47-997 |
NSP2
(VIP) |
317
aa
36,700 Da |
Nonstructural |
0 |
Orbivirus:
NS2
Orthoreovirus:
sNS |
L04531
[64] |
Non-specific
ssRNA-binding [41,62]
Accumulates in
viroplasm [65]
Involved in
viroplasm formation with NSP5 [25]
NTPase activity
[79]
Helix
destabilization activity [78]
Functional
octamer [79,80]
Binds NSP5 and
VP1 [1, 40]
Regulates NSP5
autophosphorylation [1]
Crystal
structure (HIT-like fold) [37] |
|
9
(1062) |
49-1026 |
VP7 |
326
aa
37,368 Da |
Outer
capsid glycoprotein |
780 |
- |
K02028
[4] |
Outer capsid
structural glycoprotein [21,49].
G-type
neutralization antigen [32].
N-linked high
mannose glycosylation and trimming [21].
RER
transmembrane protein, cleaved signal sequence [22].
Ca2+
binding [18].
Protection
[33]. Mediates
membrane penetration [91] |
|
10
(751) |
41-569 |
NSP4 |
175
aa
20,290 Da |
Nonstructural |
0 |
- |
AF087678
[9] |
Enterotoxin
[6].
Receptor for
budding of double-layer particle through ER membrane [5, 53].
RER
transmembrane glycoprotein [22].
Ca++/ Sr++
binding site [36].
N-linked high
mannose glycosylation [21].
Protection
[24].
Host cell [Ca2+]i
mobilization [81]. |
|
11
(667) |
22-615 |
NSP5 |
198
aa
21,725
Da |
Nonstructural |
0 |
- |
X07831
[54]
M28347
[83] |
Interacts with
VP2, NSP2 and NSP6 [1, 27].
Homomultimerizes
[27,71].
O-linked
glycosylation [28].
(Hyper-)
Phosphorylated [2, 83, 90]. Autocatalytic
kinase activity enhanced by NSP2 interaction [2].
Non-specific
ssRNA binding [52,62]. |
|
80-355 |
NSP6 |
92
aa
11,012
Da |
Nonstructural |
0 |
- |
Product of
second, out-of-frame ORF [52].
Interacts with
NSP5 [27].
Localizes to
viroplasm [52]. |
' :
Protein structure/function: RNA polymerase = A(Pol)@;
capping enzyme = A(CaP)@;
Inner virus structural protein with T = 13 symmetry = A(T13)";
viral inclusion body or viroplasm matrix protein = A(ViP)@.
Other species within the genus may have proteins with significant differences in
sizes.
†
Segments numbered based on migration of SA11 genome segments in SDS-PAGE gel.
Migration order may differ among other members of the genus.
‡ Proteins with
similar functions from other genera.
Updated April 2003, by
R.F. Ramig & M.K. Estes
(from : The RNAs and
Proteins of dsRNA Viruses: Edited by Peter. P. C. Mertens and Dennis
H. Bamford
Please make suggestions
for changes or updates to this table by e-mail to Peter Mertens peter.mertens@bbsrc.ac.uk
Reference List
1. Afrikanova, I., E. Fabbretti, M.C. Miozzo, and O.R. Burrone. 1998.
Rotavirus NSP5 phosphorylation is up-regulated by interaction with NSP2.
Journal of General Virology 79:2679-2686.
2. Afrikanova, I., M.C. Miozzo, S. Giambiagi, and O. Burrone. 1996.
Phosphorylation generates different forms of rotavirus NSP5. Journal of
General Virology 77:2059-2065.
3. Anthony, I.D., S. Bullivant, S. Dayal, A.R. Bellamy, and J.A. Berriman.
1991. Rotavirus spike structure and polypeptide composition. J.Virol.
65:4334-4340.
4.
Arias, C.F., S. L:opez, J.R. Bell, and J.H. Strauss. 1984. Primary structure
of the neutralization antigen of simian rotavirus SA11 as deduced from cDNA
sequence. J.Virol. 50:657-661.
-
Au,
K.S., W.K. Chan, J.W. Burns, and M.K. Estes. 1989. Receptor activity of
rotavirus nonstructural glycoprotein NS28. J.Virol. 63:4553-4562.
-
Ball, J.M., P. Tian, C.Q.Y. Zeng, A.P. Morris, and M.K. Estes. 1996.
Age-dependent diarrhea induced by a rotaviral nonstructural glycoprotein.
Science 272:101-104.
-
Berois, M., C. Sapin, I. Erk, D. Poncet, and J. Cohen. 2003. Rotavirus
nonstructural protein NSP5 interacts with major core protein VP2. J. Virol.
77: 1757-1763.
-
Bican, P., J. Cohen, A. Charpilienne, and R. Scherrer. 1982. Purification
and characterization of bovine rotavirus cores. J.Virol. 43:1113-1117.
-
Both, G.W., L.J. Siegman, A.R. Bellamy, and P.H. Atkinson. 1983. Coding
assignment and nucleotide sequence of simian rotavirus SA11 gene segment 10:
location of glycosylation sites suggests that the signal peptide is not
cleaved. J.Virol. 48:335-339.
-
Boyle, J.F. and K.V. Holmes. 1986. RNA-binding proteins of bovine
rotavirus. J.Virol. 58:561-568.
-
Burns, J.W., M. Siadat-Pajouh, A.A. Krishnaney, and H.B. Greenberg. 1996.
Protective affect of rotavirus VP6-specific IgA monoclonal antibodies that
lack neutralizing activity. Science 272: 104-107.
-
Chen, D., C.Q.Y. Zeng, M.J. Wentz, M. Gorziglia, M.K. Estes, and R.F.
Ramig. 1994. Template-dependent, in vitro replication of rotavirus RNA.
J.Virol. 68:7030-7039.
-
Chen, D.Y., C.L. Luongo, M.L. Nibert, and J.T. Patton. 1999. Rotavirus
open cores catalyze 5'-capping and methylation of exogenous RNA: Evidence
that VP3 is a methyltransferase. Virology 265:120-130.
-
Chizhikov V. and J.T. Patton. 2000. A four-nucleotide translation enhancer
in the 3'-terminal consensus sequence of the nonpolyadenylated mRNAs of
rotavirus. RNA 6: 814-825.
-
Clark, B. and U. Desselberger. 1988. Myristylation of rotavirus proteins.
J.Gen.Virol. 69:2681-2686.
-
Denisova, E., W. Dowling, R. LaMonica, R. Shaw, S. Scarlata, F. Ruggeri,
and E.R. Mackow. 1999. Rotavirus capsid protein VP5* permeabilizes
membranes. Journal of Virology 73:3147-3153.
-
Deo R.C., C.M. Groft, K.R. Rajashankar and S.K. Burley. 2002. Recognition
of the rotavirus mRNA 3’ consensus by an asymmetric NSP3 homodimer. Cell
106: 71-81.
-
Dormitzer, P.R. and H.B. Greenberg. 1992. Calcium chelation induces a
conformational change in recombinant herpes simplex virus-1 expressed
rotavirus VP7. Virology 189: 828-832.
-
Dormitzer P.R., Z-Y.J. Sun, G. Wagner, and S.C. Harrison. 2002. The rhesus
rotavirus VP4 sialic acid binding domain has a galectin fold with a novel
carbohydrate binding site. EMBO J. 21: 885-897.
-
Dunn, S.J., T.L. Cross, and H.B. Greenberg. 1994. Comparison of the
rotavirus nonstructural protein NSP1 (NS53) from different species by
sequence analysis and northern blot hybridization. Virology 203:178-183.
-
Ericson, B.L., D.Y. Graham, B.B. Mason, and M.K. Estes. 1982.
Identification, synthesis, and modifications of simian rotavirus SA11
polypeptides in infected cells. J.Virol. 42:825-839.
-
Ericson, B.L., D.Y. Graham, B.B. Mason, H.H. Hanssen, and M.K. Estes.
1983. Two types of glycoprotein precursors are produced by the simian
rotavirus SA11. Virology. 127:320-332.
-
Estes, M.K., D.Y. Graham, and B.B. Mason. 1981. Proteolytic enhancement of
rotavirus infectivity: molecular mechanisms. J.Virol. 39:879-888.
-
Estes, M.K., G. Kang, CQ-Y Zeng, S.E. Crawford and M. Ciarlet. 2001.
Pathogenesis of rotavirus gastroenteritis. In Gastroenteritis Viruses.
Wiley, Chichester (Novartis Foundation Symposium 238), p82-100.
-
Fabbretti, E., I. Afrikanova, F. Vascotto, and O.R. Burrone. 1999. Two
non-structural rotavirus proteins, NSP2 and NSP5, form viroplasm-like
structures in vivo. Journal of General Virology 80:333-339.
-
Fiore, L., H.B. Greenberg, and E.R. Mackow. 1991. The VP8 fragment of VP4
is the rhesus rotavirus hemagglutinin. Virology 181:553-563.
-
González, R.A., M.A. Torres-Vega, S. López, and C.F. Arias. 1998. In
vivo interactions among rotavirus nonstructural proteins. Archives of
Virology 143:981-996.
-
González, S.A. and O.R. Burrone. 1991. Rotavirus NS26 is modified by
addition of single O-linked residues of N-acetylglucosamine. Virology
182:8-16.
-
Graff J.W., D.N. Mitzel, C.M. Weisend, M.L. Flenniken and M.E. Hardy.
2002. Interferon regulatory factor 3 is a cellular partner of rotavirus
NSP1. J. Virol. 76: 9545-9550.
-
Greenberg, H.B., J. Flores, A.R. Kalica, R.G. Wyatt, and R. Jones. 1983.
Gene coding assignments for growth restriction, neutralization and subgroup
specificities of the W and DS-1 strains of human rotavirus. J.Gen.Virol.
64:313-320.
-
Groft C.M. and S.K. Burley. 2002. Recognition of eIF4G by rotavirus NSP3
reveals a basis for mRNA circularization. Mol. Cell 9: 1273-1283.
-
Hoshino, Y., M.M. Sereno, K. Midthun, J. Flores, A.Z. Kapikian, and R.M.
Chanock. 1985. Independent segregation of two antigenic specificities (VP3
and VP7) involved in neutralization of rotavirus infectivity.
Proc.Natl.Acad.Sci.U.S.A. 82:8701-8704.
-
Hoshino,Y. and A.Z. Kapikian. 1996. Classification of rotavirus VP4 and
VP7 serotypes. Arch. Virol. [Suppl] 12: 99-111.
-
Hua, J., X. Chen, and J.T. Patton. 1994. Deletion mapping of the rotavirus
metalloprotein NS53 (NSP1): The conserved cysteine-rich region is essential
for virus- specific RNA binding. Journal of Virology 68:3990-4000.
-
Hua, J., E.A. Mansell, and J.T. Patton. 1993. Comparative analysis of the
rotavirus NS53 gene: Conservation of basic and cysteine-rich regions in the
protein and possible stem-loop structures in the RNA. Virology 196:372-378.
-
Jagannath, M.R., R.R. Vethanayagam, B.S. Reddy, S. Raman, and C.D. Rao.
2000. Characterization of human symptomatic rotavirus isolates MP409 and
MP480 having 'long' RNA electropherotype and subgroup I specificity, highly
related to the P6[1],G8 type bovine rotavirus A5, from Mysore, India.
Archives of Virology 145:1339-1357.
-
Jayaram, H, Z. Taraporewala, J.T. Patton and B.V.V. Prasad. 2002.
Rotavirus protein involved in genome replication and packaging exhibits a
HIT-like fold. Nature 417: 311-315.
-
Kalica, A.R., J. Flores, and H.B. Greenberg. 1983. Identification of the
rotaviral gene that codes for hemagglutination and protease-enhanced plaque
formation. Virology. 125:194-205.
-
Kalica, A.R., H.B. Greenberg, R.G. Wyatt, J. Flores, M.M. Sereno, A.Z.
Kapikian, and R.M. Chanock. 1981. Genes of human (strain Wa) and bovine
(strain UK) rotaviruses that code for neutralization and subgroup antigens.
Virology. 112:385-390.
-
Kattoura, M.D., X. Chen, and J.T. Patton. 1994. The rotavirus RNA-binding
protein NS35 (NSP2) forms 10S multimers and interacts with the viral RNA
polymerase. Virology 202:803-813.
-
Kattoura, M.D., L.L. Clapp, and J.T. Patton. 1992. The rotavirus
nonstructural protein, NS35, possesses RNA- binding activity in vitro
and in vivo. Virology 191:698-708.
-
Kojima, K., K. Taniguchi, and N. Kobayashi. 1996. Species-specific and
interspecies relatedness of NSP1 sequences in human, porcine, bovine,
feline, and equine rotavirus strains. Archives of Virology 141:1-12.
-
LaMonica, R., S.S. Kocer, J. Nazarova, W. Dowling, E.Geimonen, R.D. Shaw
and E.R.Mackow. 2001. VP4 differentially regulates TRAF2 signalling,
disengaging JNK activation while directing NF-k B
to effect rotavirus-specific cellular responses. J. Biol. Chem. 276:
19889-19896.
-
Liu, M. and M.K. Estes. 1989. Nucleotide sequence of the simian rotavirus
SA11 genome segment 3. Nucleic.Acids.Res. 17:7991-7991.
-
Liu, M., N.M. Mattion, and M.K. Estes. 1992. Rotavirus VP3 expressed in
insect cells possesses guanylyltransferase activity. Virology 188:77-84.
-
López, S., C.F. Arias, J.R. Bell, J.H. Strauss, and R.T. Espejo. 1985.
Primary structure of the cleavage site associated with trypsin enhancement
of rotavirus SA11 infectivity. Virology. 144:11-19.
-
Ludert, J.E., N.G. Feng, J.H. Yu, R.L. Broome, Y. Hoshino, and H.B.
Greenberg. 1996. Genetic mapping indicates that VP4 is the rotavirus cell
attachment protein in vitro and in vivo. Journal of Virology 70:487-493.
-
Mansell, E.A., R.F. Ramig, and J.T. Patton. 1994. Temperature-sensitive
lesions in the capsid proteins of the rotavirus mutants tsF and tsG that
affect virion assembly. Virology 204:69-81.
-
Mason, B.B., D.Y. Graham, and M.K. Estes. 1980. In vitro transcription and
translation of simian rotavirus SA11 gene products. J.Virol. 33:1111-1121.
-
Mathieu, M., I. Petitpas, J. Navaza, J. Lepault, E. Kohli, P. Pothier,
B.V.V. Prasad, J. Cohen and F.A. Rey. 2001. Atomic structure of the major
capsid protein of rotavirus: implications for the architecture of the
virion. EMBO J. 20: 1485-1497.
-
Mattion, N.M., J. Cohen, C. Aponte, and M.K. Estes. 1992. Characterization
of an oligomerization domain and RNA-binding properties on rotavirus
nonstructural protein NS34. Virology 190:68-83.
-
Mattion, N.M., D.B. Mitchell, G.W. Both, and M.K. Estes. 1991. Expression
of rotavirus proteins encoded by alternative open reading frames of genome
segment 11. Virology 181:295-304.
-
Meyer, J.C., C.C. Bergmann, and A.R. Bellamy. 1989. Interaction of
rotavirus cores with the nonstructural glycoprotein NS28. Virology.
171:98-107.
-
Mitchell, D.B. and G.W. Both. 1988. Simian rotavirus SA11 segment 11
contains overlapping reading frames. Nucleic.Acids.Res. 16:6244-6244.
-
Mitchell, D.B. and G.W. Both. 1989. Complete nucleotide sequence of the
simian rotavirus SA11 VP4 gene. Nucleic.Acids.Res. 17:2122-2122.
-
Mitchell, D.B. and G.W. Both. 1990a. Completion of the genomic sequence of
the simian rotavirus SA11: Nucleotide sequences of segments 1, 2, and 3.
Virology 177:324-331.
-
Mitchell, D.B. and G.W. Both. 1990b. Conservation of a potential metal
binding motif despite extensive sequence diversity in the rotavirus
nonstructural protein NS53. Virology. 174:618-621.
-
Offit, P.A. and G. Blavat. 1986. Identification of the two rotavirus genes
determining neutralization specificities. J.Virol. 57:376-378.
-
Padilla-Noriega, L., O. Paniagua, S. Guzman-Leon. 2002. Rotavirus protein
NSP3 shuts off host cell protein synthesis. Virology 298: 1-7.
-
Patton, J.T. 1995. Structure and function of the rotavirus RNA-binding
proteins. Journal of General Virology 76:2633-2644.
-
Patton, J.T. 1996. Rotavirus VP1 alone specifically binds to the 3’ end
of viral mRNA, but the interaction is not sufficient to initiate
minus-strand synthesis. J. Virol. 70: 7940-7947.
-
Patton, J.T. 2001. Rotavirus RNA replication and gene expression. In Gastroenteritis
Viruses, Wiley, Chichester (Novartis Foundation Symposium 238) p64-81.
-
Patton, J.T., M.T. Jones, A.N. Kalbach, Y.W. He, and J. Xiaobo. 1997.
Rotavirus RNA polymerase requires the core shell protein to synthesize the
double-stranded RNA genome. Journal of Virology 71:9618-9626.
-
Patton, J.T., L. Salter-Cid, A. Kalbach, E.A. Mansell, and M. Kattoura.
1993. Nucleotide and amino acid sequence analysis of the rotavirus
nonstructural RNA-binding protein NS35. Virology 192:438-446.
-
Petrie, B.L., H.B. Greenberg, D.Y. Graham, and M.K. Estes. 1984.
Ultrastructural localization of rotavirus antigens using colloidal gold.
Virus.Res. 1:133-152.
-
Piron, M., T. Delaunay, J. Grosclaude, and D. Poncet. 1999. Identification
of the RNA-binding, dimerization, and eIF4GI-binding domains of rotavirus
nonstructural protein NSP3. Journal of Virology 73:5411-5421.
-
Piron, M., P. Vende, J. Cohen, and D. Poncet. 1998. Rotavirus RNA-binding
protein NSP3 interacts with eIF4GI and evicts the poly(A) binding protein
from eIF4F. EMBO Journal 17:5811-5821.
-
Pizarro, J.L., A.M. Sandino, J.M. Pizarro, J. Fernández, and E. Spencer.
1991. Characterization of rotavirus guanylyltransferase activity associated
with polypeptide VP3. J.Gen.Virol. 72:325-332.
-
Poncet, D., C. Aponte, and J. Cohen. 1993. Rotavirus protein NSP3 (NS34)
is bound to the 3' end consensus sequence of viral mRNAs in infected cells.
Journal of Virology 67:3159-3165.
-
Poncet, D., S. Laurent, and J. Cohen. 1994. Four nucleotides are the
minimal requirement for RNA recognition by rotavirus non-structural protein
NSP3. EMBO J. 13:4165-4173.
-
Poncet, D., P. Lindenbaum, R. L'Haridon, and J. Cohen. 1997. In vivo and
in vitro phosphorylation of rotavirus NSP5 correlates with its localization
in viroplasms. Journal of Virology 71:34-41.
-
Prasad, B.V., G.J. Wang, J.P. Clerx, and W. Chiu. 1988. Three-dimensional
structure of rotavirus. J.Mol.Biol. 199:269-275.
-
Prasad, B.V.V., R. Rothnagel, C.Q.Y. Zeng, J. Jakana, J.A. Lawton, W.
Chiu, and M.K. Estes. 1996. Visualization of transcriptional complexes in
rotavirus. Nature 382: 471-473.
-
Roseto, A., J. Escaig, E. Delain, J. Cohen, and R. Scherrer. 1979.
Structure of rotaviruses as studied by the freeze-drying technique.
Virology. 98:471-475.
-
Ruggeri, F.M. and H.B. Greenberg. 1991. Antibodies to the trypsin cleavage
peptide VP8* neutralize rotavirus by inhibiting binding of
virions to target cells in culture. J.Virol. 65:2211-2219.
-
Smith, R.E., S.E. Kister, and N.B. Carozzi. 1989. Cloning and expression
of the major inner capsid protein of SA-11 simian rotavirus in Escherichia
coli. Gene. 79:239-248.
-
Taniguchi, K., T. Urasawa, and S. Urasawa. 1994. Species specificity and
interspecies relatedness in VP4 genotypes demonstrated by VP4 sequence
analysis of equine, feline, and canine rotavirus strains. Virology
200:390-400.
-
Taraporewala, Z.F., J.T. Patton. 2001. Identification and characterization
of the helix-destabilizing activity of rotavirus nonstructural protein NSP2.
J. Virol. 75: 4519-4527.
-
Taraporewala, Z., D.Y. Chen, and J.T. Patton. 1999. Multimers formed by
the rotavirus nonstructural protein NSP2 bind to RNA and have nucleoside
triphosphatase activity. Journal of Virology 73:9934-9943.
-
Taraporewala Z.F., P. Schuck, R.F. Ramig and J.T. Patton. 2002. Analysis
of a rotavirus temperature-sensitive mutant indicates that NSP2 octamers are
the functional form of the protein. Journal of Virology, 76: 7082-7093.
-
Tian, P., Y. Hu, W.P. Schilling, D.A. Lindsay, J. Eiden and M.K. Estes.
1994. The nonstructural glycoprotein of rotavirus affects intracellular
calcium levels. J. Virol. 68: 251-257.
-
Vende, P., M. Piron, N. Castagné, and D. Poncet. 2000. Efficient
translation of rotavirus mRNA requires simultaneous interaction of NSP3 with
the eukaryotic translation initiation factor eIF4G and the mRNA 3' end.
Journal of Virology 74:7064-7071.
-
Welch, S.K., S.E. Crawford, and M.K. Estes. 1989. Rotavirus SA11 genome
segment 11 protein is a nonstructural phosphoprotein. J.Virol. 63:3974-3982.
-
Yang, K.J., S.X. Wang, K.O. Chang, S. Lu, L.J. Saif, H.B. Greenberg, J.P.
Brinker and J.E. Herrmann. 2001. Immune responses and protection obtained
with rotavirus VP6 DNA vaccines given by intramuscular injection. Vaccine
19: 3285-3291.
-
Zárate, S., R. Espinosa, P. Romero, E. Méndez, C.F. Arias, and S.
López. 2000. The VP5 domain of VP4 can mediate attachment of rotaviruses to
cells. Journal of Virology 74:593-599.
-
Zeng, C.Q.Y., M. Labbe, J. Cohen, B.V.V. Prasad, D. Chen, R.F. Ramig and
M.K. Estes. 1994. Characterization of rotavirus VP2 particles. Virology 201:
55-65.
-
Zeng, C.Q.Y., M.J. Wentz, J. Cohen, M.K. Estes, and R.F. Ramig. 1996.
Characterization and replicase activity of double-layered and single-layered
rotavirus-like particles expressed from baculovirus recombinants. Journal of
Virology 70:2736-2742.
-
Prasad,
B.V.V. Burns, J.W., Marietta, E., Estes, M.K., and Chiu, W. 1990,
Localization of VP4 neutralization sites in rotavirus by three-dimensional
electron cryo-microscopy, Nature, 343: 476-479.
-
Shaw,
A., L., Rothnagel, R., Chen, D., Ramig, R.F., Chiu, W. and Prasad, B.V.V.
1993
Three-dimensional visualization of rotavirus hemagglutinin structure.
Cell 74:693-701.
-
Blackhall, J., M. Munoz, A. Fuentes, and G. Magnusson. 1998.
Analysis of rotavirus non-structural protein NSP5 phosphorylation.
J. Virol. 72: 6398-6405.
-
Charpilienne A, Abad MJ, Michelangeli F, Alvarado F, Vasseur M, Cohen J,
Ruiz MC. (1997)
Solubilized and cleaved VP7, the outer glycoprotein of rotavirus, induces
permeabilization of cell membrane vesicles. J Gen Virol.,
78:
1367-71.

|